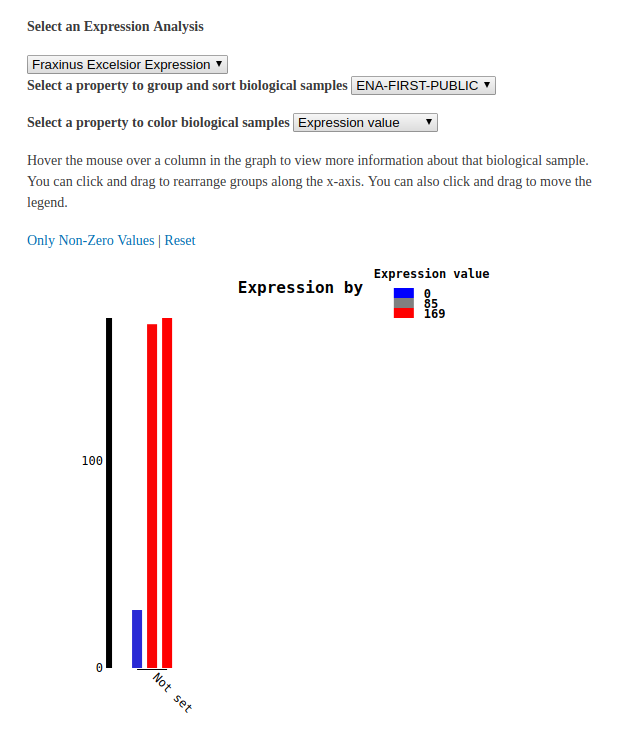
Loading Expression Data¶
Create Analysis¶
You will first need an analysis to associate the expression data. To do so, navigate to Tripal_content -> Add Tripal Content. Select Analysis.
- Name - Something along the lines of Fraxinus Excelsior Expression.
- Program, Pipeline, Workflow or Method Name - Something along the lines of e.g DESESQ2.
- Program Version - Something along the lines of e.g 1.0.
- Date Performed - Leave at default.
Load the expression data¶
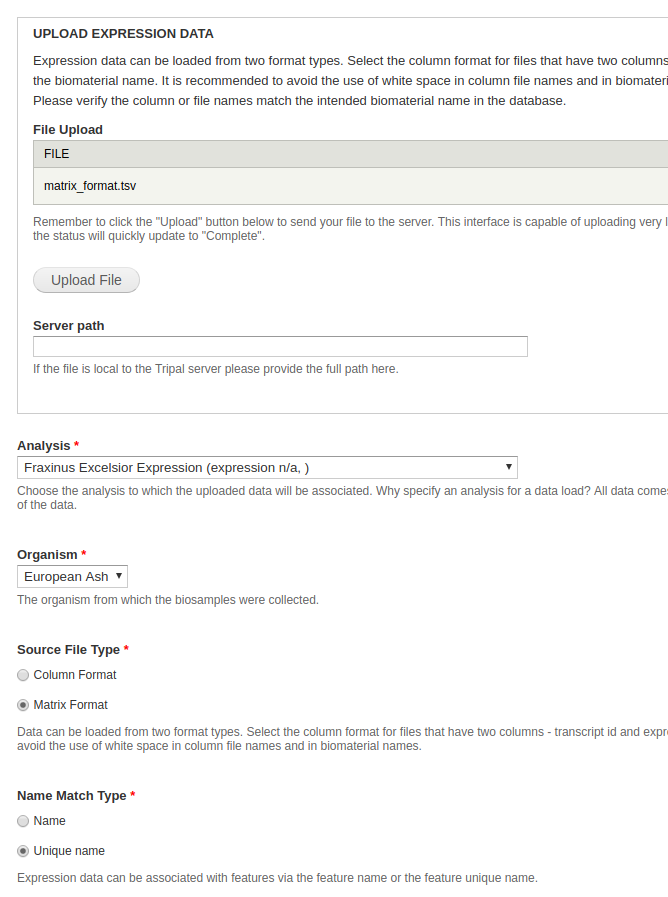
Expression data is loaded by the tripal_analysis_expression module using the Chado Expression loader, located at admin/tripal/loaders/Chado_Expression_Data_Loader. Expression data should be in column or matrix format.
- File Upload - For this dataset, the simplest method is uploading the .tsv file.
- Analysis - Select the same analysis specified for the biosamples.
- Organism - Select an organism. In this case, the organism is European Ash.
- Source File Type - If you’re uploading the .tsv file, select Matrix. If you’re uploaded the .txt files, select Column. Keep in mind that if you upload the .tsv file, you do not need to upload the .txt files.
- Name Match Type - Select unique name.
All other fields can be ignored. Click Import expression data. A green header should appear at the top of the page with a job for you to run. Run it and you’re done.
Publishing¶
Publishing is not necessary for expression data, as we don’t create any new Tripal Entities.
Viewing Expression Results¶
The easiest way to check to see if your expression results were successfully uploaded is by referring to the chado_elementresult table. If the table has contents, you know the results were uploaded successfully.
If you can’t access the database, the alternative is to display the expression results directly on a feature page. Results are hidden by default, so we have to enable them in order to view them. This can be done with the admin menu by navigating to Structure > Tripal Content Types. In the row mRNA, click Manage Fields and towards the top of the window, click Check for new fields. This will take a moment, but a new field should be found called data__gene_expression_data.
At the top of the window, click Manage Display. Scroll all the way to the bottom of the window and look for a Disabled field, in which an Expression field should be contained. Move this out of the disabled table.
Now our results should be available to view. Navigate to any feature page (from the admin menu, Content > Tripal Content, click any record with type mRNA) and you should see your expression results.